lesionBrain
lesionBrain is a pipeline to automatically segment white matter lesions from MRI data (T1w + FLAIR). It gets anonymized MRI brain volumes in NIFTI format and produces a pdf report with the volumes of the lesions and their locations.
Labelling protocol
White matter lesions were manually outlined by an expert radiologist using multimodal MRI data (T1w + FLAIR) to create a library of 43 cases. After the segmentation process, lesions are classified based on their location as periventricular, deep white, juxtacortical and infratentorial.
Report
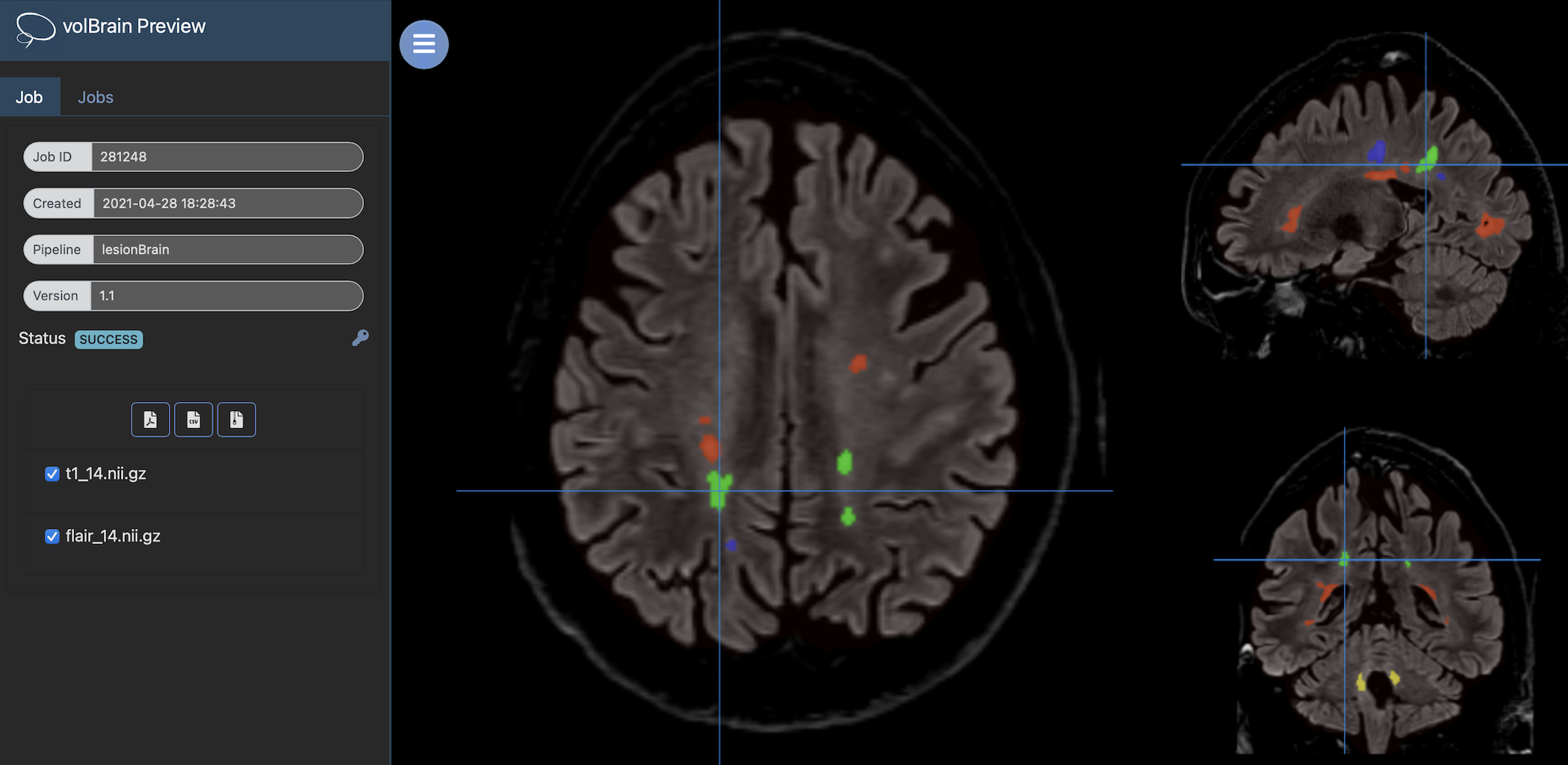
Once the process is finished, you will be notified by e-mail so you will be able to download a package including some image files and two reports (CSV and PDF) offering all the volumes estimated from the segmentations. As you can see in the figure below, the PDF includes patient information, lesion types, their volumes and locations in the MNI space. It also includes several snapshots from the different labeling steps as a quality control.
Download PDF Report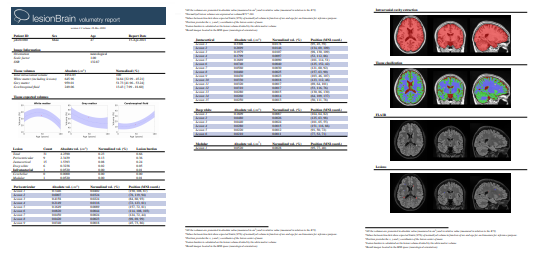
Version 1.0: This version is based on our 43 training images and does not include patch-based error correction (see reference below).
Version 1.1: This version is based on our 43 training images as well as additional images manually segmented by volbrain users who have shared them with us, thanks to them ;-). This version includes all the steps described in the original paper (see reference below).
References
Pierrick Coupé, T. Tourdias, P. Linck, Jose E. Romero, Jose V. Manjón. Lesionbrain:an online tool for white matter lesion segmentation. International Workshop on Patch-based Techniques in Medical Imaging, 95-103, 2018. PDF