AssemblyNet
AssemblyNet is a pipeline of processes aimed to automatically analyze MRI brain data. It gets an anonymized MRI brain volume in NIFTI format and produces a pdf report with the volumes of all the brain structures.
Labelling protocol
All the segmentations are based on the Neuromorphometrics protocol which comprises 132 structures. In this protocol, the segmentation of subcortical structures follows the "general segmentation" protocol as defined by the MGH Center for Morphometric Analysis. Moreover, the segmentation of cortical structures follows the BrainCOLOR protocol. In AssemblyNet, the brain structures are fused to produce cortex lobes and brain tissues.
AssemblyNet can used as a standalone application by downloading its docker here.
Report
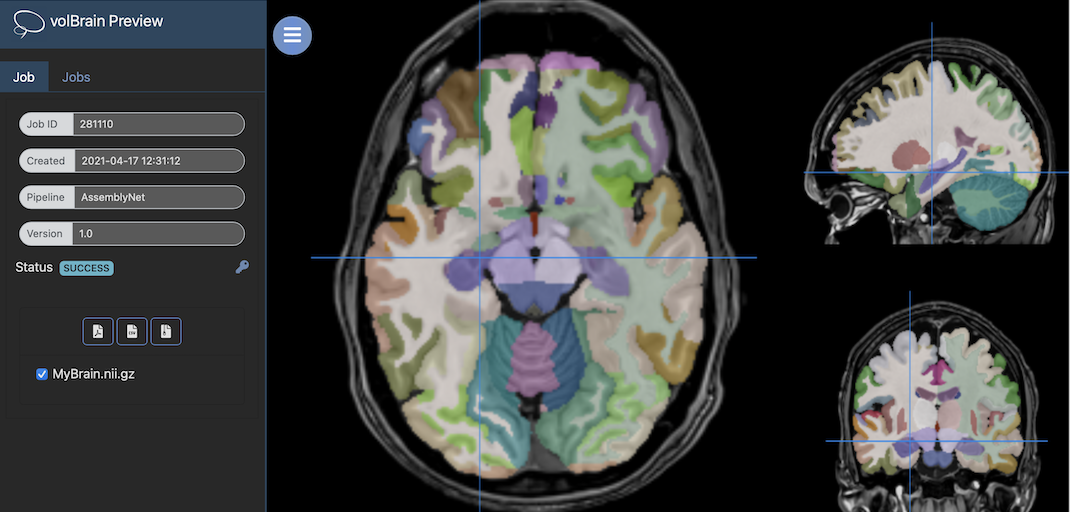
Once the process is finished you will be notified by e-mail so you will be able to download a package including some image files and two (CSV and PDF) reports gathering all the volumetry values calculated from the segmentations. As you can see in the figure below, the PDF includes patient information, the volumes of parenchyma, brain tissues, macrostructures and 132 brain structures as well as asymmetry indexes. Finally, it also includes several snapshots from the different labeling steps as a quality control.
Download PDF Report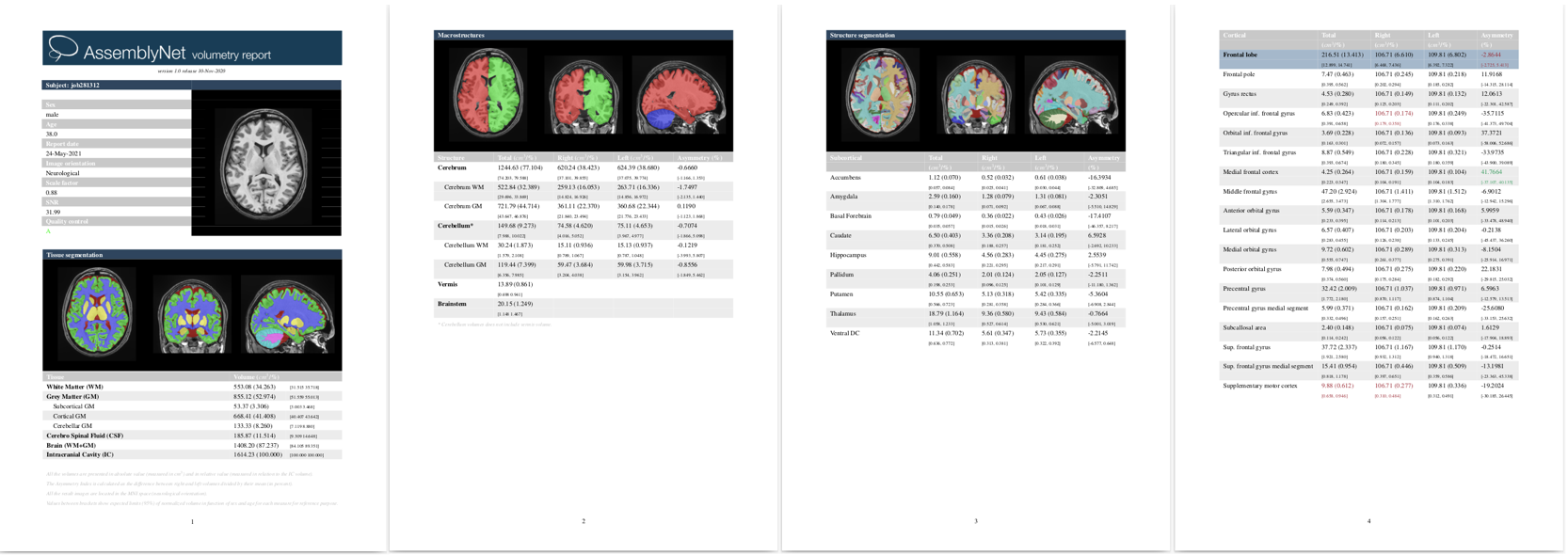
References
P. Coupé, B. Mansencal, M. Clément, R. Giraud, B. Denis de Senneville, V.-T Ta, V. Lepetit, J. V. Manjon. AssemblyNet: A large ensemble of CNNs for 3D Whole Brain MRI Segmentation. NeuroImage, 219, 117026, 2020. PDF
de Senneville, B.D., Manjon, J.V. and Coupé, P., 2020. RegQCNET: Deep quality control for image-to-template brain MRI affine registration. Physics in Medicine & Biology, 65(22), p.225022. PDF